Microscopy Nodes
Microscopy Nodes
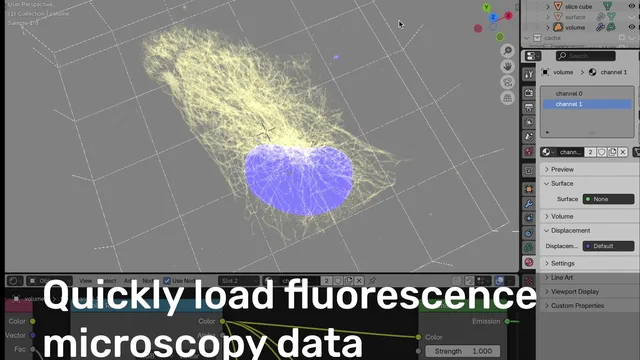
Microscopy Nodes helps load microscopy data and visualize it for easy, accurate, and intuitive data visualization of microscopy!
Currently, Microscopy Nodes supports the import of up to 5D tif files (xyz, time and channels), with segmentation channels loaded as meshes instead of volumes. Microscopy Nodes allows the user to set pixel size in µm, and provides auto-generated editable scale boxes to annotate volume data.
In short, Microscopy Nodes supports:
- Up to 5D .tif files (tzcyx, in any order)
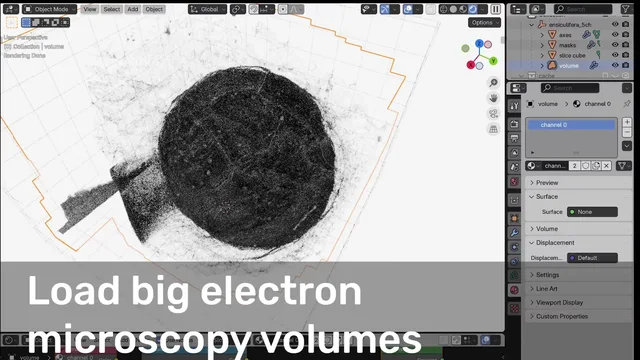
- Adapted to handle both fluorescence and electron microscopy
- Loading segmentation masks as meshes
- Automatic µm-scale axes
- Arbitrary slicing, multiple slicing objects
- Auto-loaded isosurface renders
Microscopy Nodes takes arbitrary sizes of image arrays, as long as this fits within RAM and manages to resave into the blender-native .vdb file format.
Microscopy Nodes used to be known as 'Tif2Blender'. Tutorials for Microscopy Nodes can be found on https://www.youtube.com/@oanegros
Usage
The Microscopy Nodes panel can be found in Scene Properties:
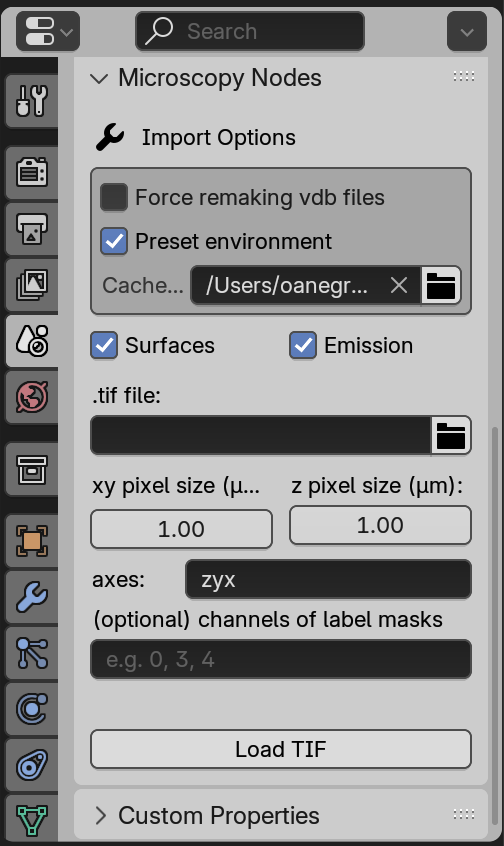
Here we have multiple parameters that can be set, most notably:
.tif file- the tif microscopy file to load
Cache directory- where local files will be stored (remember to clear this out once in a while)
The addon should set the pixel sizes and axis order automatically for imagej-tif files, but you can always edit them in the panel.
For electron microscopy data:
- it is recommended to turn off
Surfaces(does not load an isosurface mesh, which may take too much RAM) andEmission(takes the volumetric data as absorbing light rather than emitting light).
label masks:
- The
label masksinterface allows you to note which channels contain integer-identity label masks, and these will be loaded as meshes. Note that for binary masks, theSurfacesmay be more effective.
hyperparameters:
Force remaking vdb files- Microscopy Nodes will overwrite any existing files with this filename in the Cache directory
Preset environment- Microscopy Nodes default-environment variables that make it possible to render huge volumes in relatively short times.
What's New
1.1.4 August 12th, 2024
Remove reliance on Blender initial names:
- 'World' name for presetting env
- move all MiN objects to new collection Microscopy Nodes to not prefer a 'Collection' instance being present so that this does not accidentally becomes essential
Activity
-
-
For a reviewer, i have sample data for a tutorial of an outdated version of this plugin at https://github.com/oanegros/Blender_for_Biologists_2023/tree/main/data (published under CC-BY)
-
- nickberckley changed review status to Awaiting Changes
- 1 mo
There is assets folder which seems to include just another zip of add-on? It creates nesting, should be removed.
I don't think you need to package numpy module, it is shipped with Blender. Same can be true for others, I suggest you research this a bit.
Can you provide what is the difference between windows, linux and macos versions in the code?
-
This add-on uses
str.strip,str.lstriporstr.rstrip, taking multiple characters that seem to assume order.All characters are stripped in any order.
So
"test_nojs.json".strip(".json")results in"test_", nottest_nojs.Use
str.removeprefix&str.removesuffixinstead.Ref:
.strip('obj')&.strip('ch'). -
Thanks for the reviews @nickberckley and @campbell, new version 1.3 no longer has nesting or installs numpy, and changes these instances of
str.striptostr.removeprefixfor clarity. The other dependencies (i use two packages,scikit-imageandtifffilewhich have some subsequent depencencies) are all necessary and not shipped with Blender. The OS version split exists because bothscikit-imageand its dependencyscipyhave platform-dependent builds. Otherwise there is no difference between the platform versions in the code, and i've split the versions to not bloat the zip files (i was struggling to upload a previous version with all wheels of all platforms). -
@Oane-Gros typically
str.strip/str.removeprefixis not just about clarity, strip may remove characters in any order which may have unexpected results (e.g. ).strip('obj')will strip.jobas well as.obj.
Checking further this extension:
This add-on makes error prone assumptions about data block names in the users file: e.g.
bpy.data.worlds["World"]The active world should be used instead.
Ref:
TifLoadOperatorcalls topreset_environment&preset_em_environment. -
Hi, thanks again for reviewing this!
@campbell on the point of
str.strip: here the strings that are stripped were created above and the behaviour was thus relatively determinate, butstr.removeprefixwill make it easier to maintain as the behaviour is more specific to the usage :)I've removed the
bpy.data.worlds["World"]from the preset env functions, in the same vein, the add-on used to prefer a base collection called"Collection"(however not having this was not breaking), this assumption on Blender default name was removed as well. -
You have to comment with "Awaiting Review" tag to prompt moderators if you're ready for another round of review.
-
-
-
-
Hey everyone,
Just a heads-up about a recent change regarding the licensing of add-ons on the Blender extension platform. Moving forward, all add-ons will need to be released under the GNU/GPL 3.0 license (SPDX:GPL-3.0-or-later). This is mainly to keep things simple and consistent across the board.
Previously, we accepted various licenses as long as they were compatible with Blender’s distribution. However, to avoid any confusion and streamline the process, all add-ons using the bpy API should now be presented as GPL 3 (the same license the Blender bundle is distributed). Regardless of whether the original code was under GPL 2, or something else like MIT or ZLIB.
Existing add-ons versions won't be affected. However, new updates will need to comply to the revised requirements.
Thanks for understanding, and feel free to reach out if you have any questions.
-
-
- nickberckley changed review status to Awaiting Changes
- 2 w
I tested loading tif file that you have on your github (tests), and I don't understand why is performance that bad. Looking at geonodes and shader setups its nothing extreme, very simple ones, but even in solid mode it is very bad performance. What is causing that?
On extensions side I suggest you provide either screenshots with location of add-on or a brief description about where it is and how to use it. Its not apparent now without going on github. I would also suggest closing the panel on default so it doesn't clutter space when add-on isn't used, but that's on you
-
Hi @nickberkley, I'll work on the description on the extensions page, good point, I'm still used to most of my users coming from github/youtube.
For the performance, I think this is mostly due to the fact that this is still quite a large volume at 1024x1024x195 = 204 MP even though this is already undersampled from the microscope, as microscopy data can get up to TBs. The resulting surface is 17 M triangles. I indeed specifically keep the geonodes and shaders simple to be able to handle big volumes. I'm working on a major update that will allow setting surface resolution on load and also to load smaller versions of data if the file type is pyramidal, but this will still take some time and it would be nice for the microscopy community if the current version could already be on the extension platform.
-
- nickberckley changed review status to Approved
- 2 w
Approving with expectation that description will be improved
-
-
-
-
-
-
-
-
-
Add comment
Sign in to comment.



Ready for review